SomaRestingVmTest¶
Validate Resting membrane voltage from soma of Purkinje cell¶
-
class
cerebunit.validation_tests.cells.Purkinje.test_for_soma_restVm.SomaRestingVmTest(*args, **kwargs)¶ This test compares the measured resting Vm observed in real animal (in-vitro or in-vivo, depending on the data) generated from neuron against those by the model.
The test class has three levels of mechanisms.
Level-1
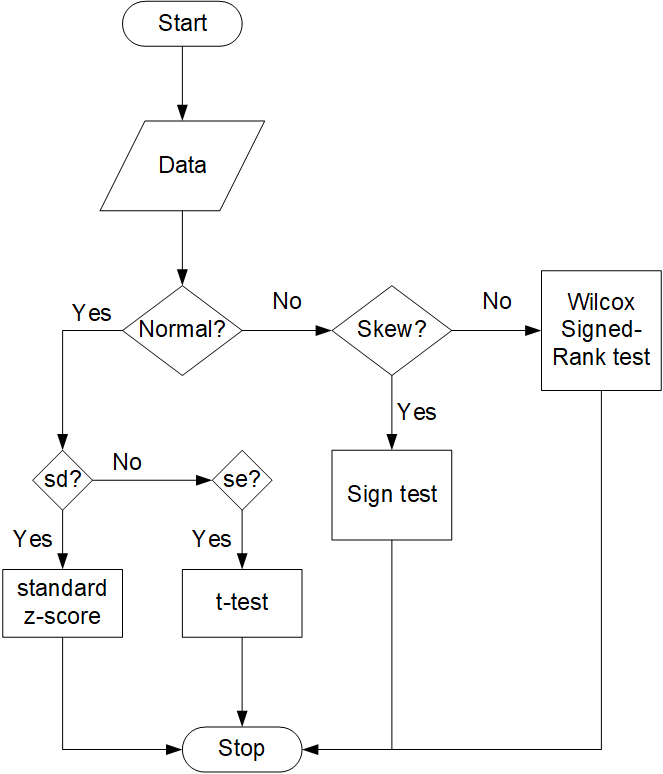
validate_observation()Given that the experimental/observed data has the following: mean, SD (or SE), sample_size, units, and raw_data,
validate_observation()checks for them. The method then checks the data condition by askingNecessaryForHTMeans. Depending on the data condition the appropriatescore_typeis assigned and corresponding necessary parameter; for t-Test, the parameterobservation["standard_error"]and for sign-Test, the parameterobservation["median"].Level-2
generate_prediction()The model is executed to get the model prediction. The prediction is a the resting Vm from the soma of a PurkinjeCell returned as a
quantities.Quantityobject.Level-3
compute_score()The prediction made by the model is then used as the null value for the compatible
score_typebased on the data condition (normality and skewness) determined byvalidate_observation(). The level ends by returning the compatible test-statistic (t or z-statistic) as ascore.How to use:
from cerebunit.validation_tests.cells.Purkinje import SomaRestingVmTest data = json.load(open("/home/main-dev/cerebdata/expdata/cells/PurkinjeCell/Llinas_Sugimori_1980_soma_restVm.json")) test = SomaRestingVmTest( data ) s = test.judge(chosenmodel, deep_error=True)
Then to get the test score
s.scoreand test report callprint(s.description). If one is interested in getting the computed statistics calls.statistics.Further notes on the test.
The experimental observation data (as json file) must have the element protocol_parameters, which in turn has the nests the elements temperature and initial_resting_Vm.
One should consider whether the model is compared against in vitro or in vivo experimental data (in addition to the species under study). For example,
- Consider the Llinas and Sugimori (1980, 10.1113/jphysiol.1980.sp013357) experimental data (Llinas_Sugimori_1980_soma_restVm.json)
- The reported experimental data only includes those with initial resting levels for \(\geq -50 mV\) discarding those for \(< -50 mV\).
- The observed resting potential are claimed by the authors to be more negative than those observed in vivo.
- The authors infer that this could be due to in vitro which is done on slices. The slicing removes background synaptic input generated by parallel fibre synapses.
- For more details see Llinas_Sugimori_1980_soma_restVm.json
-
compute_score(observation, prediction, verbose=False)¶ This function like generate_pediction is called automatically by sciunit which RestingVmTest is a child of. This function must be named compute_score The prediction processed from “vm_soma” is compared against the experimental_data to get the binary score; 0 if the prediction correspond with experiment, else 1.
-
generate_prediction(model, verbose=False)¶ Generates resting Vm from soma. The function is automatically called by sciunit.Test which this test is a child of. Therefore as part of sciunit generate_prediction is mandatory.
-
validate_observation(observation, first_try=True)¶ This function is called automatically by sciunit and clones it into self.observation This checks if the experimental_data is of some desired form or magnitude. Not exactly this function but a version of this is already performed by the ValidationTestLibrary.get_validation_test